Our Research
Synthetic Microbiology
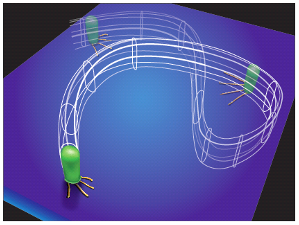 Our research in this area focuses on developing a unique microbial tracking system that
enables us to monitor entire bacterial communities with single-cell resolution
over multiple generations. Through this system, we study surface sensing,
multicellular behavior coordination, and multigenerational memory transfer,
which could provide insights into how bacteria interact with their environment
and each other, with potential implications for medicine, agriculture, and
environmental science.
Our research in this area focuses on developing a unique microbial tracking system that
enables us to monitor entire bacterial communities with single-cell resolution
over multiple generations. Through this system, we study surface sensing,
multicellular behavior coordination, and multigenerational memory transfer,
which could provide insights into how bacteria interact with their environment
and each other, with potential implications for medicine, agriculture, and
environmental science.
Innate Immune Peptides and Antimicrobials
 Our goal is to gain a deeper understanding of the sequences of innate immune
peptides, and therefore structure and function, through the application of
machine learning, geometry, and coordination chemistry. We are currently
investigating the relationships between autoimmunity and innate immunity with
various bodily systems, including the nervous, cardiovascular, digestive, and
endocrine systems. This includes exploring the properties of host antimicrobial
peptides (AMPs), antimicrobial neuropeptides for CNS defense, cell-penetrating
peptides, mitochondrial remodeling proteins, apoptosis proteins, enterotoxins,
and other related molecules.
Our goal is to gain a deeper understanding of the sequences of innate immune
peptides, and therefore structure and function, through the application of
machine learning, geometry, and coordination chemistry. We are currently
investigating the relationships between autoimmunity and innate immunity with
various bodily systems, including the nervous, cardiovascular, digestive, and
endocrine systems. This includes exploring the properties of host antimicrobial
peptides (AMPs), antimicrobial neuropeptides for CNS defense, cell-penetrating
peptides, mitochondrial remodeling proteins, apoptosis proteins, enterotoxins,
and other related molecules.
- Innate immunity and antimicrobial peptides
- Antibiotic-resistant pathogens and antibiotic design
- Transporter sequences and cell penetrating peptides for drug delivery
- Budding and scission peptides in viruses
- Autoimmune diseases (Ex: lupus, psoriasis)
Synthetic Immunology
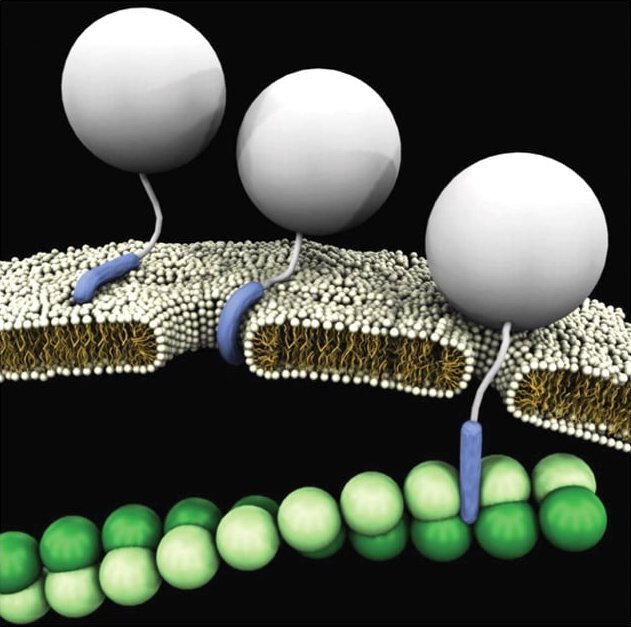
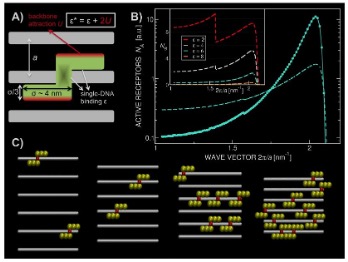 We have found that innate immune peptides hold a
remarkable ability to guide and organize Toll Like Receptor (TLR) ligands into
crystalline complexes that can present themselves in a multivalent manner. These
complexes are spaced in such a way that corresponds to the size of TLRs,
resulting in a heightened immune response. We have observed this phenomenon in
plasmacytoid dendritic cells with dsDNA/TLR9 and in keratinocytes with
dsRNA/TLR3. Our team has also found that these supramolecular complexes can be
programmed to assemble in a manner that is either pro-inflammatory or
anti-inflammatory, making them a promising avenue for further research in
immunology.
We have found that innate immune peptides hold a
remarkable ability to guide and organize Toll Like Receptor (TLR) ligands into
crystalline complexes that can present themselves in a multivalent manner. These
complexes are spaced in such a way that corresponds to the size of TLRs,
resulting in a heightened immune response. We have observed this phenomenon in
plasmacytoid dendritic cells with dsDNA/TLR9 and in keratinocytes with
dsRNA/TLR3. Our team has also found that these supramolecular complexes can be
programmed to assemble in a manner that is either pro-inflammatory or
anti-inflammatory, making them a promising avenue for further research in
immunology.
The Physical Immunology of COVID-19
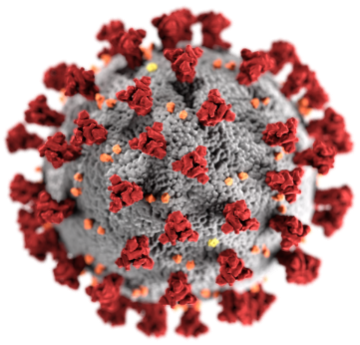 We are deeply interested in studying the physical factors that make the
coronavirus responsible for COVID-19 so hyperinfectious and hyperinflammatory.
More specifically, we seek to understand how SARS-CoV-2 induces cytokine storms,
catastrophic coagulation, skin rashes and lesions (MIS-C, 'covid fingers and
toes'), as well as the connection between COVID and autoimmune diseases such as
lupus, rheumatoid arthritis, and atherosclerosis. Additionally, we aim to
investigate why some coronaviruses are capable of inducing these consequences
while others are not and the phenomenon of 'long COVID'.
We are deeply interested in studying the physical factors that make the
coronavirus responsible for COVID-19 so hyperinfectious and hyperinflammatory.
More specifically, we seek to understand how SARS-CoV-2 induces cytokine storms,
catastrophic coagulation, skin rashes and lesions (MIS-C, 'covid fingers and
toes'), as well as the connection between COVID and autoimmune diseases such as
lupus, rheumatoid arthritis, and atherosclerosis. Additionally, we aim to
investigate why some coronaviruses are capable of inducing these consequences
while others are not and the phenomenon of 'long COVID'.
Bacterial Communities and Biofilms
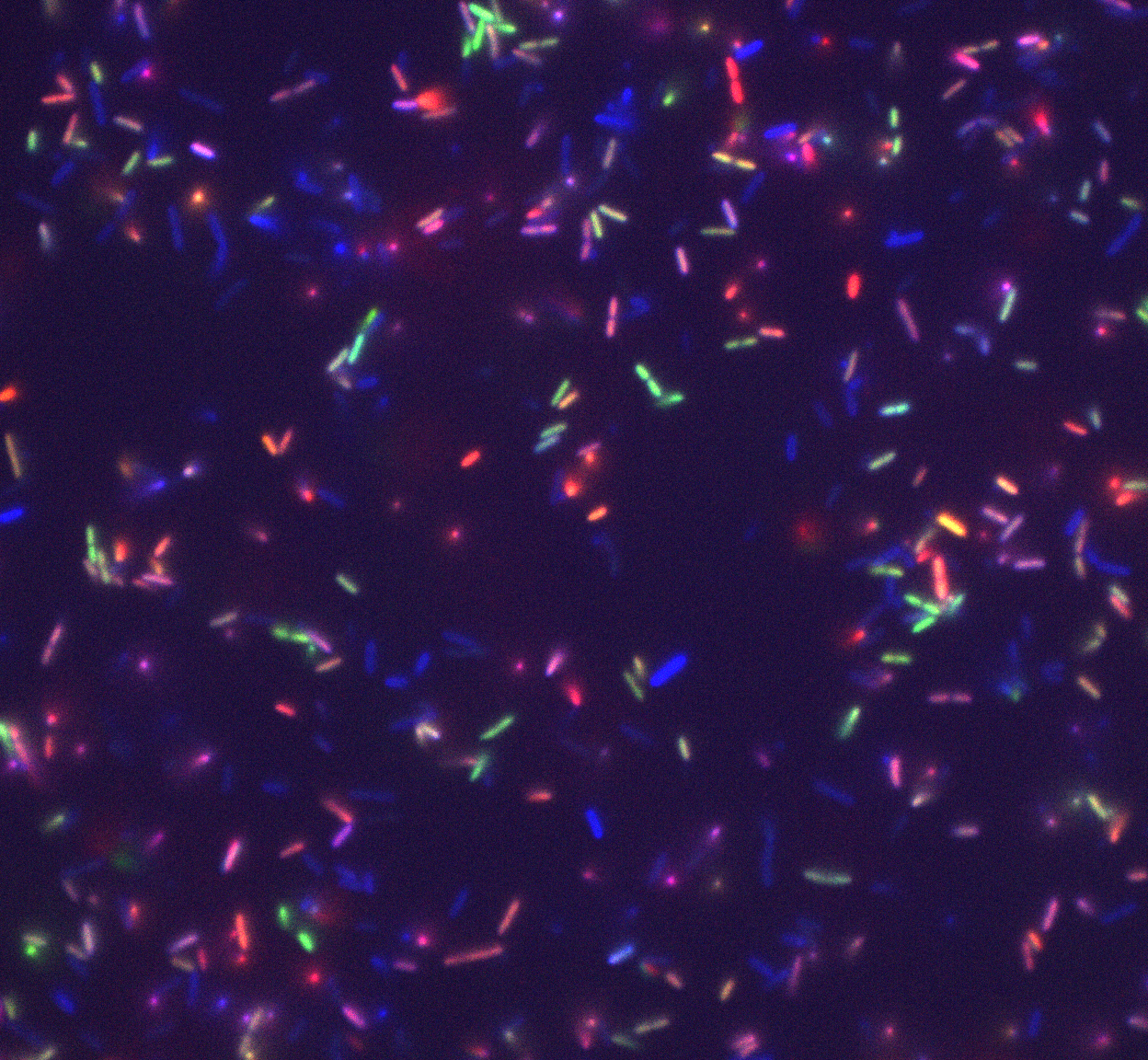 We are interested in the progression of a biofilm from its inception to mature
state and subsequent dispersal of new progenitor cells. The first step in forming
a biofilm is a single cell making contact with a surface. One of the questions
that we study is "How does a cell know that it is on a surface?". We utilize time-
lapse microscopy of flow cell systems and image processing to track the development
of a biofilm from its initial cell, analyzing its behavior and expression as
it divides and progresses through the early stages of biofilm formation. Our research
has implications for infections biofilm protects bacteria from antibiotic intervention
making treatment difficult, such as in implants and in cystic fibrosis (CF).
Some of our areas of research include:
We are interested in the progression of a biofilm from its inception to mature
state and subsequent dispersal of new progenitor cells. The first step in forming
a biofilm is a single cell making contact with a surface. One of the questions
that we study is "How does a cell know that it is on a surface?". We utilize time-
lapse microscopy of flow cell systems and image processing to track the development
of a biofilm from its initial cell, analyzing its behavior and expression as
it divides and progresses through the early stages of biofilm formation. Our research
has implications for infections biofilm protects bacteria from antibiotic intervention
making treatment difficult, such as in implants and in cystic fibrosis (CF).
Some of our areas of research include:
- Bacterial Communities and Interactions
- cdGMP and cAMP Signalling
- Cellular Surface Motility
- Bacterial Swarming
- Social Organization in Biofilms
- Image processing, machine learning, application of 'big data' methods
- Biofouling
- Cystic Fibrosis
Enabling Fundamental Science: Physics, Chemistry, Biology
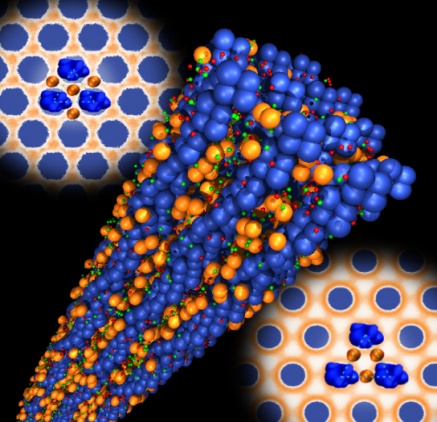
- Protein-lipid interactions and membrane curvature generation (Ex: Apoptosis proteins and cancer therapeutics)
- Synchrotron x-ray methods
- Physical chemistry of solvation (Ex: femtosecond hydration dynamics of ions and surfaces)
- Soft condensed matter physics of self-assembly (Ex: polymers, polyelectrolytes, membranes, liquid crystals, colloids, nanoparticles)